Findings can open the way to broad-spectrum antiviral drugs against ZIKV
Though initially described as a mild version of dengue fever, the Zika virus (ZIKV) outbreak in the Americas unexpectedly revealed major neurological impacts as fetal microcephaly or other congenital brain injuries when women are infected during pregnancy and Guillain-Barre´ syndrome (a disorder in which the body’s immune system damages the nerves) in adults. It can be transmitted both by the insect vector and sexual contact. Its outbreak became a global health threat of complex epidemiology and devastating neurological impacts, therefore requiring urgent efforts towards the development of novel efficacious and safe antiviral drugs.
A prime target for drug discovery is the non-structural protein 5 (NS5) RNA-dependent RNA-polymerase (RdRp) which is encoded in the genomes of all RNA-containing viruses with no DNA stage and has a central role in RNA viral replication
Although nucleoside polymerase inhibitors (NPIs) have achieved clinical success in the case of Hepatitis C virus infections (for example, sofosbovir), they depend on activation by host kinases and are potentially subjected to toxicity problems. Therefore, non-NPIs have been actively sought as inhibitors of flaviviral NS5 RdRp, particularly targeting the so-called priming loop, that regulates RNA-template binding and polymerization. Recently, several groups reported the discovery of novel RdRp inhibitors with pan-serotype activity against dengue viruses (DENV). In these cases, the use of X-ray crystallographic structures was fundamental to develop optimized lead candidates [1, 2].
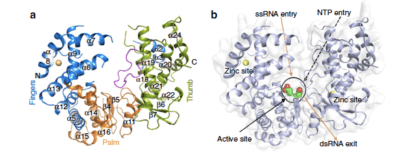
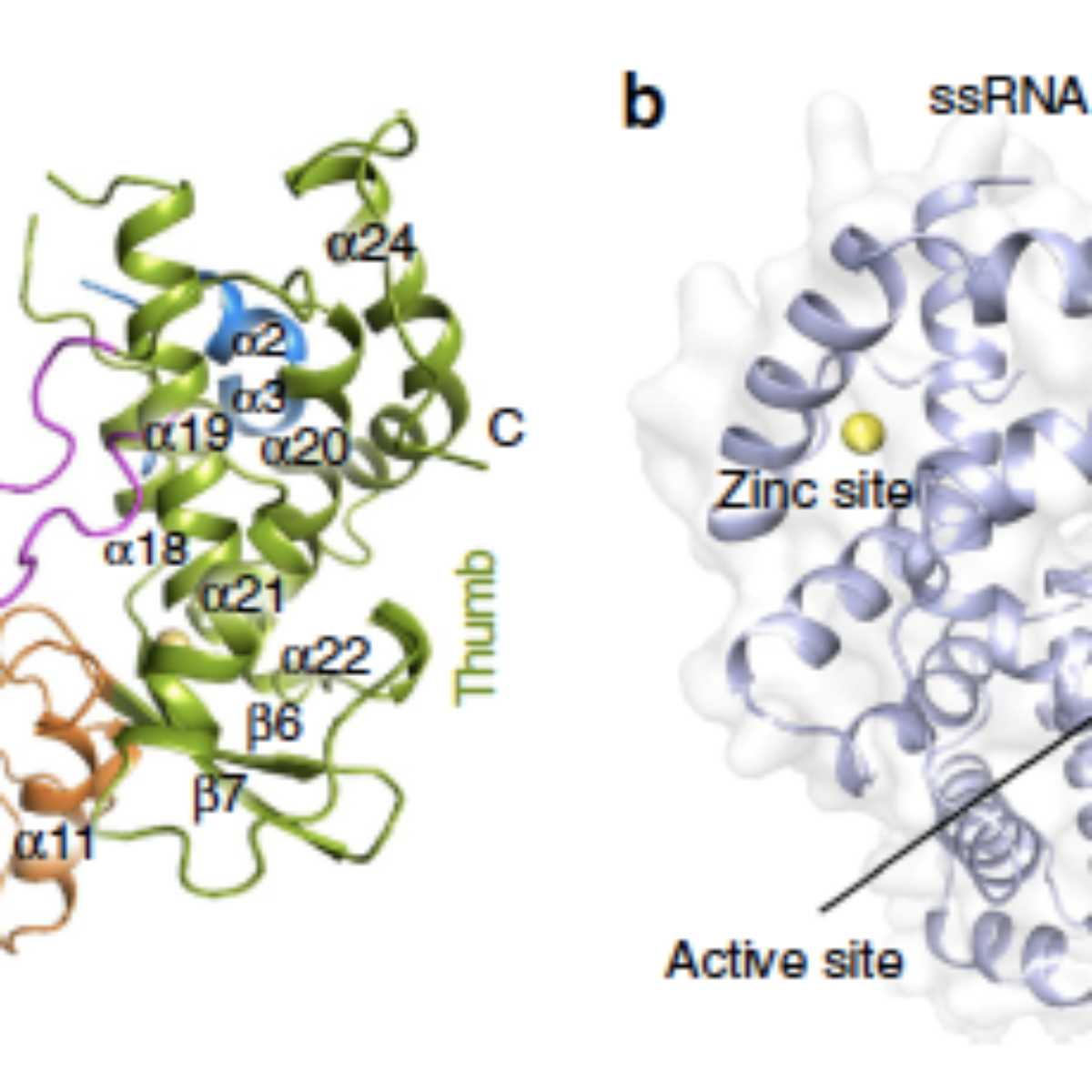
Figure 1. Crystal structure of Zika virus NS5 RdRp. (a) ZIKV NS5 RdRp structure with fingers, palm and thumb domains coloured in blue, orange and green, respectively. The priming loop is depicted in pink. (b) Surface view of the ZIKV NS5 RdRp, with orange arrows pointing the entry of the single-strand RNA template, and the exit region of the double-strand RNA. Catalytic aspartates are depicted as green/red spheres. Black dashed arrow shows the entry path of NTPs, while black arrow point to the position of the active site. In both figures, zinc atoms are depicted in yellow.
Recently, Andre S. Godoy et al. [3], have determined the crystal structure of the ZIKV NS5 RdRp domain at high resolution (1.9 Å) and have compared it with the homologous dengue virus proteins from different serotypes to identify suitable target sites for anti-ZIKV drug discovery and elucidate their structural drug-binding features. The overall structure is similar to other flaviviral homologue (dengue, West Nile virus, yellow fever virus….). However, the priming loop target site, which is suitable for non-nucleoside polymerase inhibitor including a tighter pocket and a modified local charge distribution.
Single crystals were cryo-cooled in liquid nitrogen and X-ray diffraction data were collected at the beam line MX2 of the Brazilian Synchrotron Light Laboratory (LNLS) using a wavelength of 1.4586 Å. Despite the structural similarities with DENV, the ZIKV NS5 RdRp exhibits significant differences in the priming loop binding site that may affect ligand design.
These findings can open the way to the discovery and development of broad-spectrum antiviral drug candidates against Flaviviridae, including ZIKV.
Sources
[1] Yokokawa, F. et al. Discovery of potent non-nucleoside inhibitors ofdengue viral RNA-dependent RNA polymerase from a fragment hit using structure-based drug design. J. Med. Chem. 59, 3935–3952 (2016). DOI: 10.1021/acs.jmedchem.6b00143
[2] Anusuya, S., Velmurugan, D. & Gromiha, M. M. Identification of dengue viral RNA-dependent RNA polymerase inhibitor using computational fragmentbased approaches and molecular dynamics study. J. Biomol. Struct. Dyn. 34,1512–1532 (2016). DOI: 10.1080/07391102.2015.1081620
[3] Andre S. Godoy, Gustavo M.A. Lima, Ketllyn I.Z. Oliveira, Naiara U. Torres, Fernando V. Maluf, Rafael V.C. Guido & Glaucius Oliva. Crystal structure of Zika virus NS5 RNA-dependent RNA polymerase. Nature Communications 8, 14764, 2017. DOI:10.1038/ncomms14764
Researchers investigate the preservation of soft tissues during fossilization
Reaction is an important step in the transformation of carbon dioxide into fuels